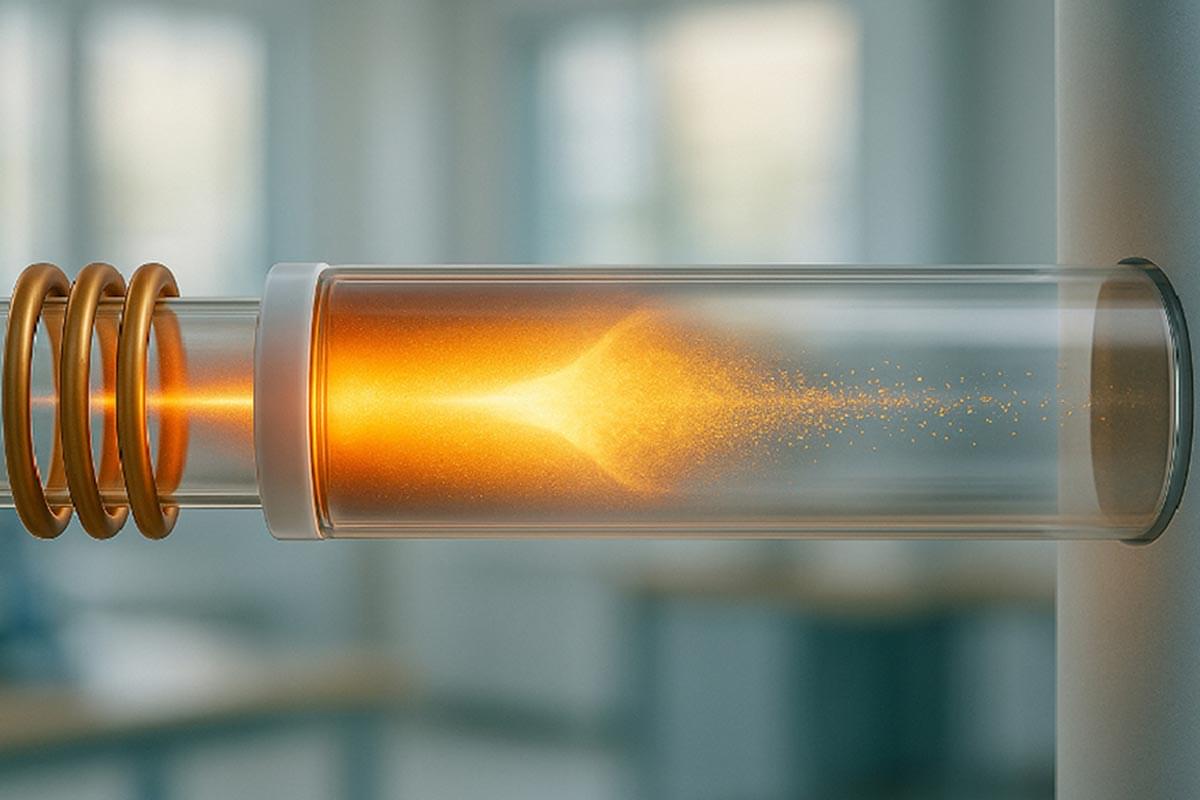
A breakthrough depolymerization technology is transforming plastic waste into their base chemical building blocks, offering a scalable alternative to traditional recycling with strong unit economics.



KU Leuven, Belgium bioscience engineers have developed a roadmap, so to speak, for industrial cellulose gasoline.
The bioscience engineers already knew how to make gasoline in the laboratory from plant waste such as sawdust. In 2014, at KU Leuven’s Centre for Surface Chemistry and Catalysis, the researchers succeeded in converting sawdust into building blocks for gasoline.
A chemical process made it possible to convert the cellulose – the main component of plant fibers – in the sawdust into hydrocarbon chains. These hydrocarbons can be used as an additive in gasoline. The resulting cellulose gasoline is a second-generation biofuel.
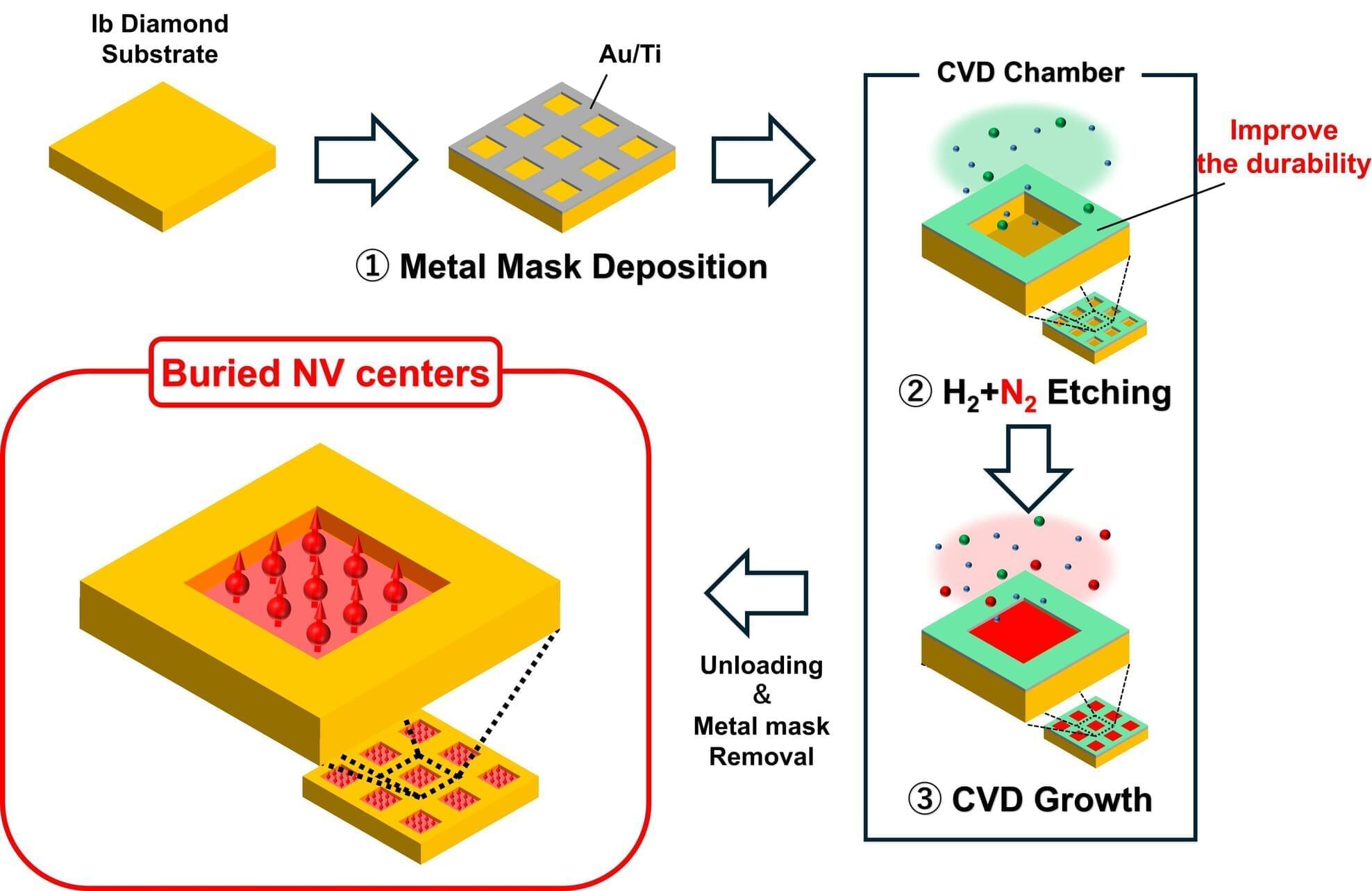
Researchers at Kanazawa University, in collaboration with Diamond and Carbon Applications (Germany), have developed a buried-growth process for nitrogen–vacancy (NV) centers in diamond using microwave plasma chemical vapor deposition (MPCVD). By employing nitrogen-radical selective etching, which simultaneously enhances metal-mask durability through nitridation, the team enabled a continuous etching–growth sequence within a single MPCVD process.
The work is published in the journal Carbon.
Optical measurements confirmed highly aligned NV centers selectively buried in predefined regions. This integrated approach provides a stable and scalable platform for orientation-controlled diamond qubits and future room-temperature quantum technologies.
Can pure water store electrical energy? A research team led by Dr. Vasily Artemov within the Cluster of Excellence “BlueMat—Water-Driven Materials” at Hamburg University of Technology has now shown that it can. By confining water within nanometer-sized channels in clay minerals, the researchers created a supercapacitor capable of efficiently storing and transporting electrical charge.
What makes the finding unusual is that it uses pure water as its electrolyte—the medium that transports electrical charge. Today’s batteries and supercapacitors typically rely on added salts, acids, or other chemical electrolytes. In contrast, the new system works without such additives and is based solely on abundant, naturally occurring materials: water, clay, and carbon.
“Our goal is to develop safer and more sustainable energy-storage technologies based on abundant materials rather than complex chemical compounds,” says Artemov, lead author of the paper published in Nature Communications. “The device stores and releases energy efficiently, operates at a comparatively high voltage for a water-based system, and remains stable over tens of thousands of charging cycles.”

While Mythos 5 remains largely unconstrained for restricted government and trusted enterprise partners, Fable 5 is wrapped in a sophisticated safety perimeter. If Fable 5 detects a prompt drifting toward high-risk vectors—like cyberwarfare exploits, advanced biology, or chemical synthesis—it doesn’t just give a generic “I can’t answer that” error. Instead, the query seamlessly falls back to Claude Opus 4.8 (Anthropic’s next-most capable model) to handle the response safely.
Today we’re launching Claude Fable 5: a Mythos-class1 model that we’ve made safe for general use.
Fable 5’s capabilities exceed those of any model we’ve ever made generally available. It is state-of-the-art on nearly all tested benchmarks of AI capability, showing exceptional performance in software engineering, knowledge work, vision, scientific research, and many other areas. The longer and more complex the task, the larger Fable 5’s lead over our other models.
Releasing a model this capable comes with risks. Without safeguards, Fable 5’s capabilities in areas like cybersecurity could be misused to cause serious damage. We’ve therefore launched the model with safeguards that mean queries on some topics will instead receive a response from our next-most-capable model, Claude Opus 4.8. To release the model both safely and quickly, we’ve tuned these safeguards conservatively—they’ll sometimes catch harmless requests, though they trigger, on average, in less than 5% of sessions. With more capable models arriving in the coming months, we’re working to improve our safeguards and reduce false positives as quickly as we can.
The energy-efficient desalination system produces fresh water without chemical additives and transforms leftover salts into useful materials.
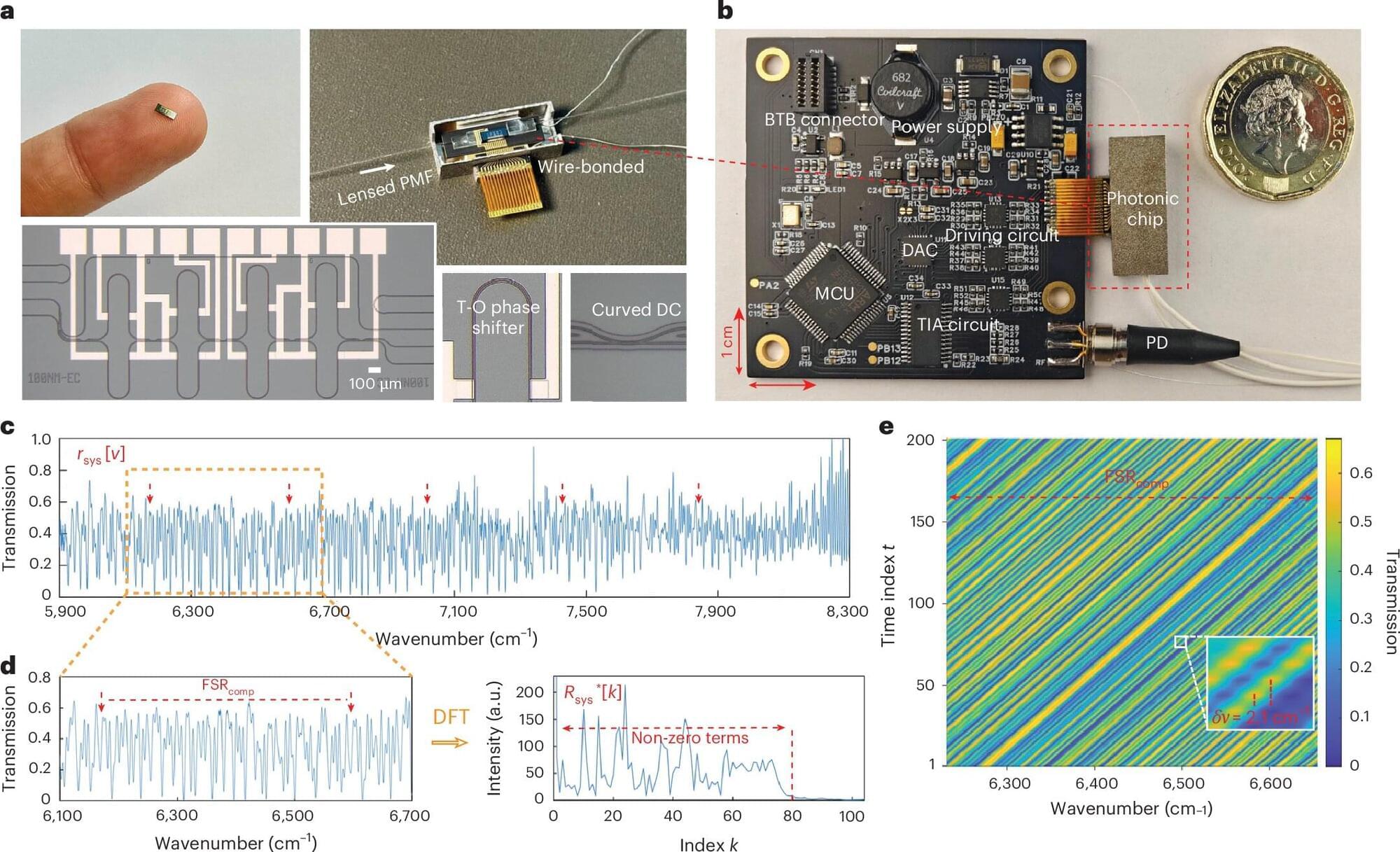
Researchers from the University of Cambridge and GlitterinTech, a startup founded by the same research group, have unveiled a fundamentally new type of optical spectrometer that delivers laboratory-grade precision in a device small enough to be embedded in portable and wearable technologies. By rethinking how spectra are measured and processed, the team has demonstrated a spectrometer costing only around $10, operating at a centimeter scale, and capable of applications ranging from industrial quality control to real-time health care monitoring.
Optical spectrometers underpin countless technologies, from chemical analysis and manufacturing to environmental sensing and medicine. Yet shrinking these instruments has historically involved painful trade-offs: Miniaturized devices typically sacrifice bandwidth, resolution or accuracy, limiting them to rough identification rather than true metrological measurements. The newly reported convolutional spectrometer overcomes these barriers by introducing a conceptually elegant operating principle grounded in the convolution theorem, offering unprecedented performance metrics compared with existing dispersive, Fourier-transform and reconstructive spectrometers.
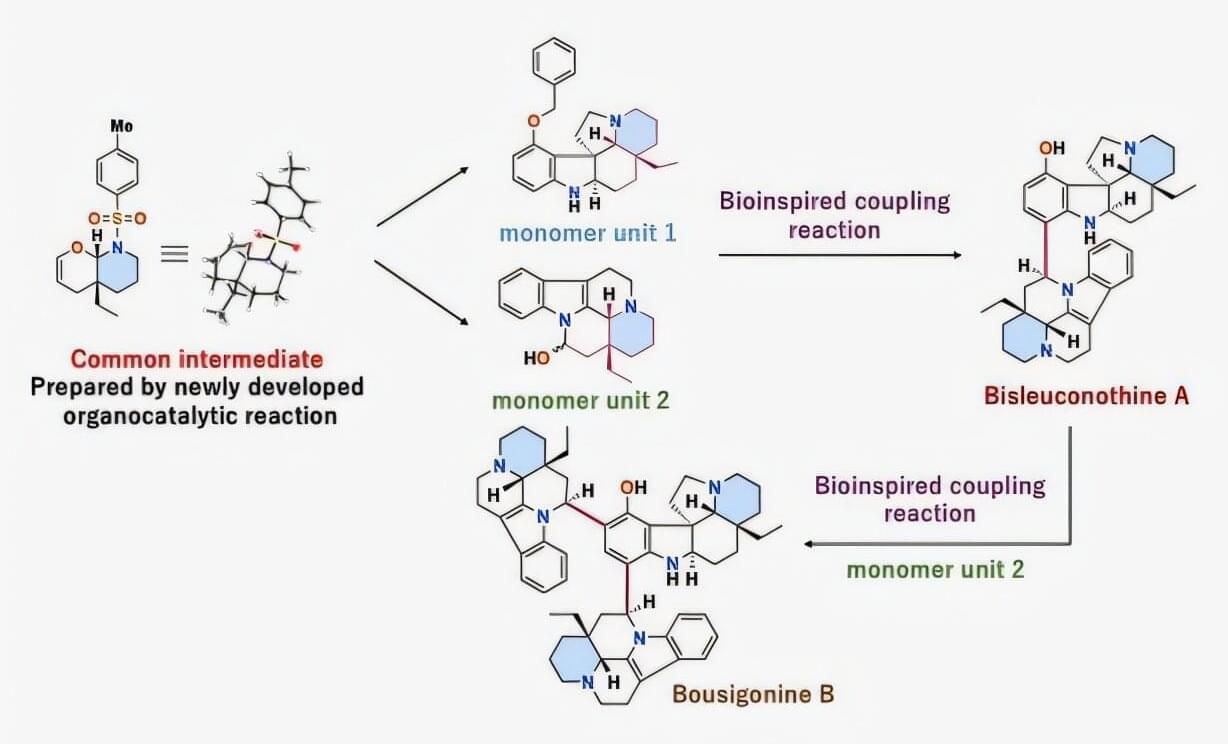
Plants are undeniably one of nature’s most promising sources of new medicines, with monoterpenoid indole alkaloids (MIAs) being a great example. Some intricate compounds are built from multiple-linked chemical units that form highly complex three-dimensional structures. Because of their size and shape, scientists believe such oligomeric MIAs may be able to interfere with specific protein–protein interactions inside cells—a biological target that conventional small-molecule drugs often struggle to reach.
This unusual capability could make MIAs uniquely suited to combat various diseases. Such is the case for bisleuconothine A, an MIA isolated from plant bark in 2010 that has shown strong activity against breast cancer and lung cancer.
Despite their therapeutic potential, these compounds are extremely difficult to produce synthetically in the laboratory. Their structures contain multiple interconnected rings and several precisely arranged stereocenters, meaning their atoms must be assembled in the correct three-dimensional orientation to preserve their biological activity. Because of this, drug development research involving oligomeric MIAs remains limited.

A new study finds that ancient hominins nearly 800,000 years ago deliberately selected specific basalt sources for different stages of tool production rather than simply using whatever stone was available nearby. By tracing the geochemical “fingerprints” of stone tools to both exposed and now-buried basalt flows, the researchers demonstrated that these hominins possessed detailed environmental knowledge, advanced planning abilities, and long-term technological traditions that were maintained and repeated across generations.
A new study published in Scientific Reports provides new insights into the technological behavior and raw material procurement strategies of early Middle Pleistocene hominins at the Acheulian site of Gesher Benot Ya’aqov (GBY). The study uses geochemical analyses of basalt artifacts and nearby basalt sources to trace where the raw material used for tool production came from and to reconstruct how early hominins selected stone within a landscape that has changed dramatically over time. The research was carried out by Dr. Tzahi Golan and Dr. Yoav Ben Dor of the Geological Survey of Israel, and Prof. Naama Goren-Inbar of the Hebrew University of Jerusalem.
GBY, dated to about 780,000 years ago, preserves repeated occupations of Acheulian hominins along the shores of paleo-Lake Hula. Excavations directed by Prof. Goren-Inbar revealed a rich archaeological record, including stone tools made of flint, limestone and basalt, as well as evidence of fire use, plant exploitation, animal processing and fish consumption.
