Nature is filled with remarkable visual phenomena created by microscopic surface structures that interact with light in fascinating ways. The iridescent wings of butterflies, the shimmering feathers of birds and the glossy surfaces of flower petals are all examples of how living organisms control the reflection, absorption and scattering of light. These optical effects are not only visually striking but also serve important biological functions, including attracting pollinators, communication, camouflage and protection from environmental stress. Understanding these naturally occurring photonic structures has become an important area of research, as they provide inspiration for the development of advanced biomimetic materials and optical technologies.
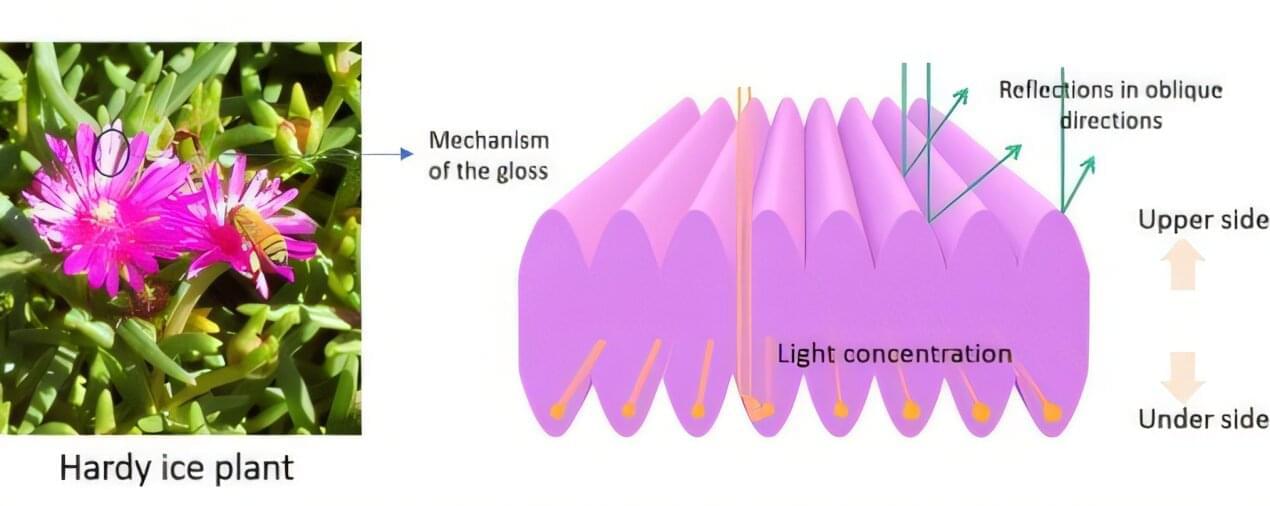
One such example is the hardy ice plant, Delosperma cooperi, a perennial succulent native to South Africa and widely cultivated in Japan. The flower’s petals display a striking glossy appearance, prompting researchers to investigate the mechanism responsible for this effect.
Researchers from Shinshu University, led by professor Hiroshi Moriwaki, conducted this study to understand how the petals generate gloss and whether their surface structure could inspire the design of novel reflective materials. Kazuma Tanabe also was part of the research team. The findings are published in the journal Optical Materials.