Researchers relied on a newer gene-editing technique that may make it possible to engineer embryos, a prospect that has long alarmed bioethicists.

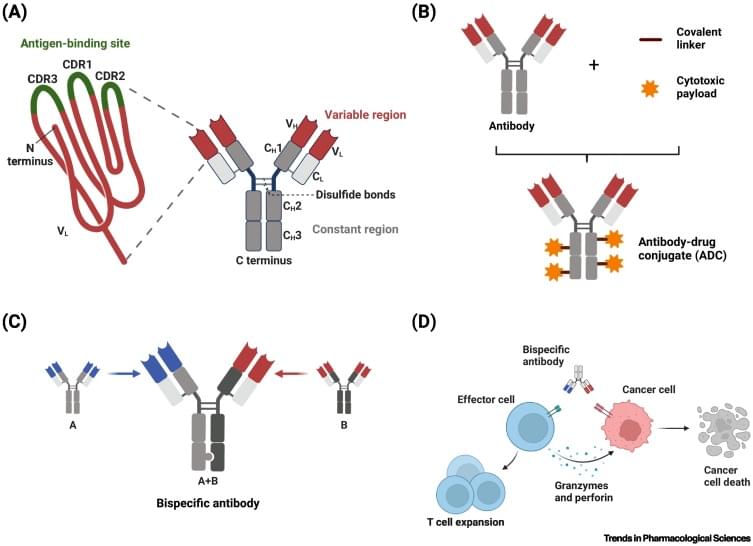
Antibodies in oncology are being equipped with toxic cargoes and effector functions that can kill cells at very low concentrations. A key challenge is that most targets on cancer cells are also present on at least some healthy cells. Shared targets can result in off-tumor binding and compromise the safety and potential of therapeutic candidates. In this review, we survey strategies that can help direct biologics to cancer sites more selectively. These strategies are becoming increasingly feasible thanks to advances in molecular design and engineering. The objective is to create therapeutics that exploit changes in cancer and leverage the human body infrastructure, enabling therapeutics that discriminate not just self from non-self but diseased from healthy tissue.
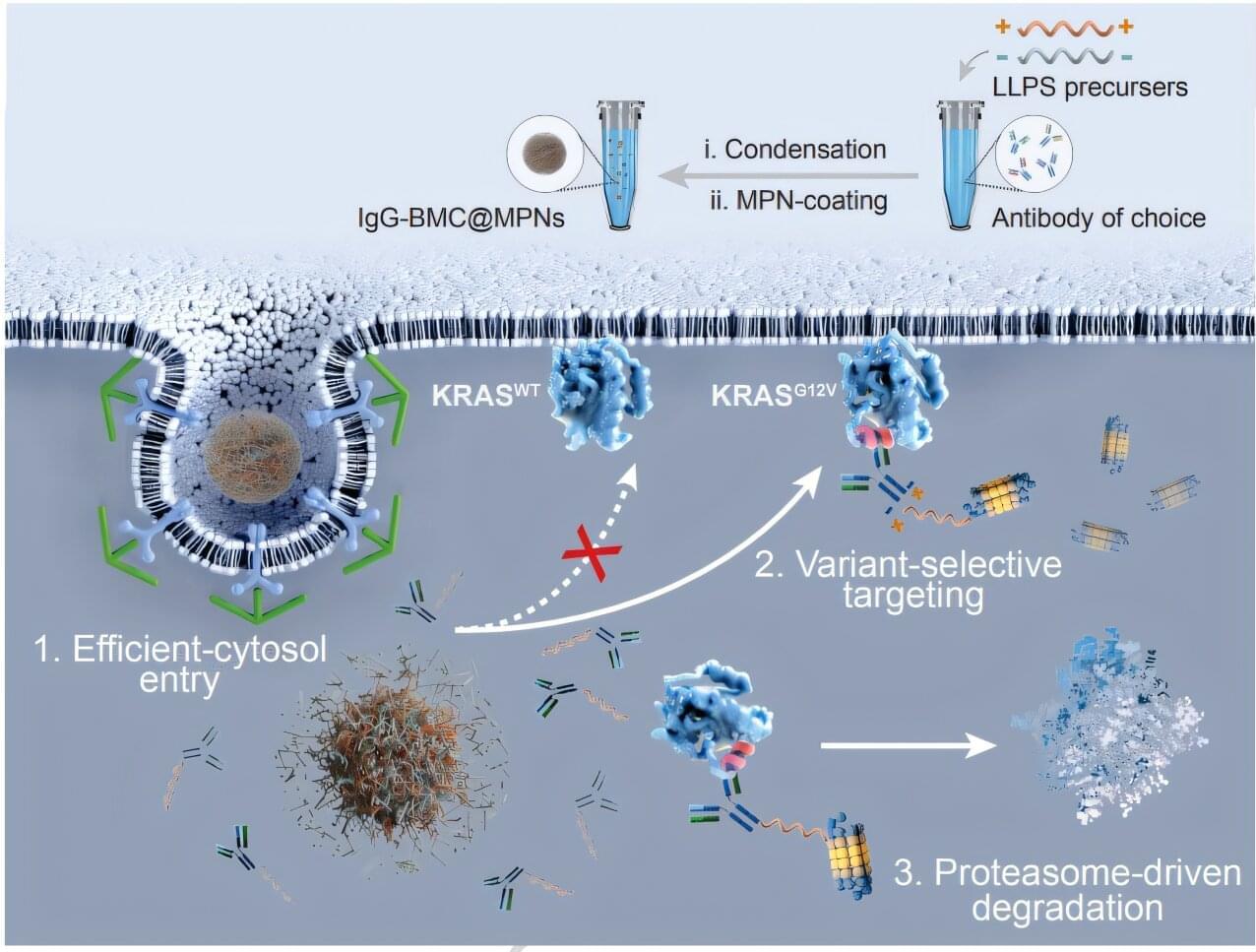
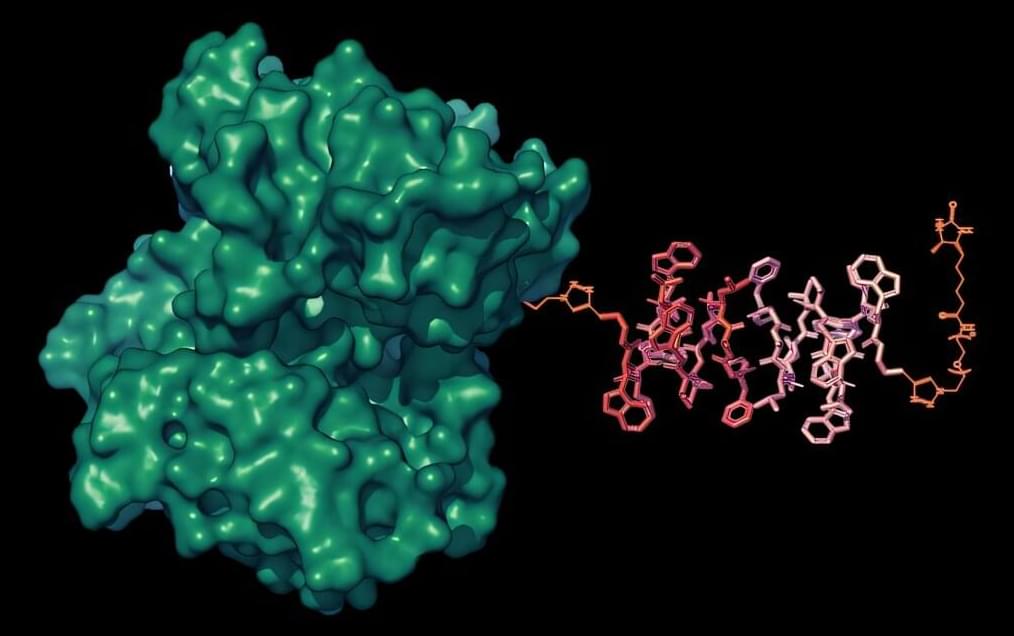
Northwestern Medicine scientists have developed a novel synthetic biomolecular condensate that can degrade intracellular disease-causing proteins, providing a framework for new therapeutic approaches for a wide range of diseases, as detailed in a recent study published in Nature Communications.
Shana Kelley, Ph.D., the Neena B. Schwartz Professor of Chemistry, Biomedical Engineering, and Biochemistry and Molecular Genetics and the president of the Chan Zuckerberg Biohub Chicago, was senior author of the study.
Targeted protein degradation is an emerging therapeutic strategy that harnesses cells’ own degradation machinery to clear disease-causing proteins. However, achieving this degradation process across different cell types has remained a challenge due to subtle variations in protein structure.
On April 21, the Munk Debates convened a special debate about gene editing in Deerfield, Massachusetts for 650 students at Deerfield Academy.
Motion: Be it Resolved, let’s engineer better human beings.
About the Debate:
New powerful engineering technology is already being used to edit human embryos, curing diseases and repairing defective genes before a child is even born. Some welcome this new science as a powerful tool to enhance human intelligence, memory, appearance and physical health. Why wouldn’t we embrace a science that allows people to live longer, healthier, and happier lives? Others warn that this new technology will be used to create designer babies and a new class of genetically “enhanced” elites. It will undermine human dignity and autonomy, and risk unleashing new diseases into the human gene pool. Playing G-d with human nature, critics argue, will result in a dystopian nightmare of our own making.
About the Debaters:
Arguing in favour of the motion was the biophysicist, best-selling author, biotechnology entrepreneur, and the former director of the Program on Medicine, Technology and Society at UCLA School of Medicine, Gregory Stock. His debate partner was the internationally acclaimed strategic philosopher and pioneering transhumanist Max More. Arguing against the motion was the prominent American bioethicist Ezekiel Emanuel, Special Advisor to the Director General of the WHO and a former founding chair of the Department of Bioethics at the NIH. His debate partner was the award-winning educator, author, and Professor of Reproductive Science at University College London, Joyce Harper.
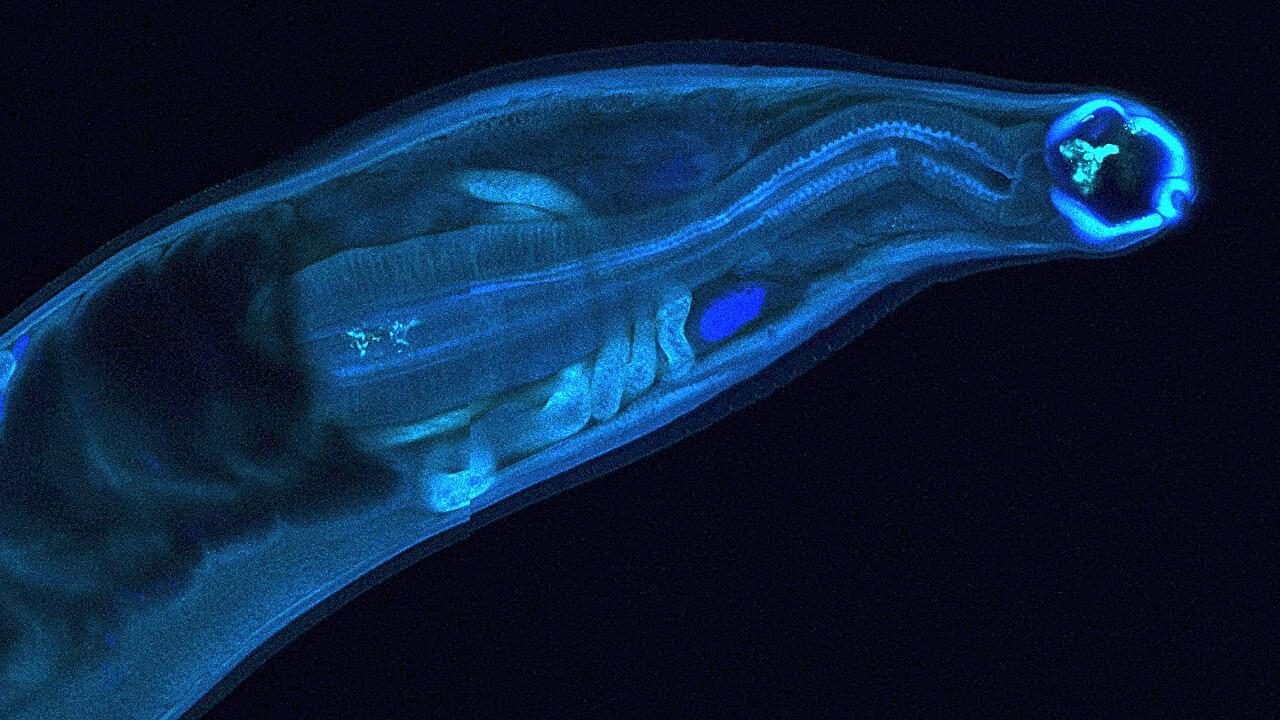
Hookworms, intestinal parasites that infect hundreds of millions of people in under-resourced tropical regions around the globe, have evolved to survive inside the human gut for years, secreting molecules that enable coexistence with their hosts. Now, researchers at Washington University School of Medicine in St. Louis have harnessed that biological mechanism for potential human benefit, engineering a hookworm to produce and deliver a drug within a living host.
In a new study, the team reports the first successful genetic modification of the human hookworm. It was designed to produce an antibody that neutralizes tetrodotoxin, a deadly neurotoxin produced by pufferfish and other marine animals. After colonizing an animal host with the modified hookworms, the parasites produced the antitoxin and secreted it into the bloodstream, partially inactivating the toxin. The findings are published in Nature Communications.
The work demonstrates that this drug production and delivery approach could be a long-term solution to any number of medical needs, from chronic conditions requiring continuous drug treatment to exposure to toxins in remote locations without medical care available.
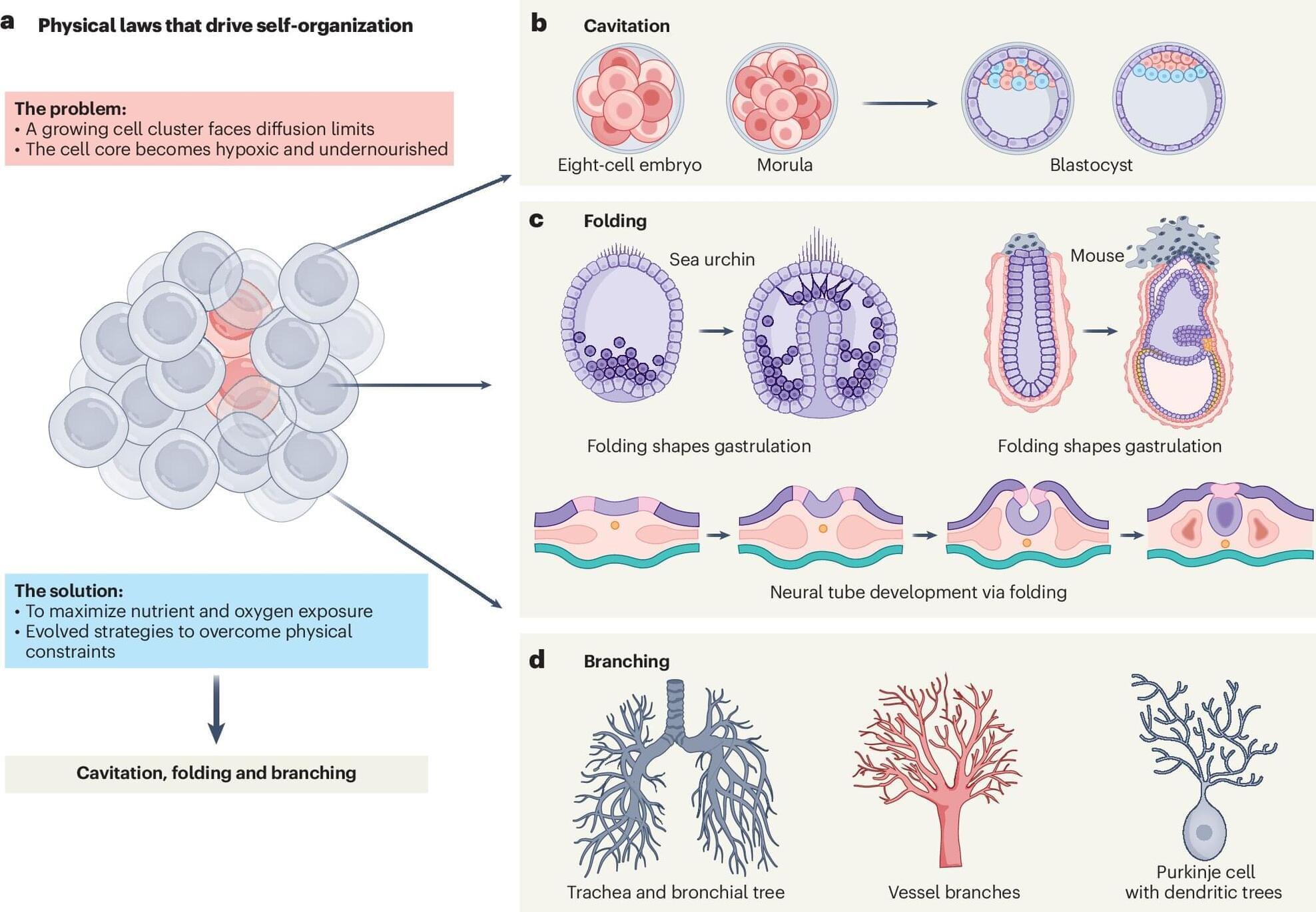
How did life make the leap from single cells to coordinated, multicellular organisms? And how do genetically identical cells still perform a version of that feat every time an embryo begins to take shape?
In a new Perspective paper appearing in the journal Nature Biotechnology, Bren Professor of Biology and Biological Engineering Magdalena Zernicka-Goetz and collaborator Qi Chen of the University of Utah ask one of biology’s oldest questions in a new way. The paper is titled “Decoding the origins of cellular self-organization for engineered biology.”
New LDL Drug Could Cure Heart Disease. Eli Lilly published Phase 1 data in the New England Journal of Medicine showing that a single IV infusion of a gene editing therapy called VERVE-102 lowered LDL cholesterol permanently. Well… the effect held for at least 18 months. Longevity Twitter immediately called it the cure for heart disease. The science is real. The hype is getting ahead of what the paper actually says. This episode walks through the Phase 1 Heart-2 trial — the data, the base-editing mechanism (which is NOT CRISPR), the one safety event nobody’s talking about, and how Eli Lilly’s CEO is publicly thinking about pricing a one-and-done cure.
HUME BODY POD DISCOUNT UP TO 50% OFF:
Code: LSN20
https://humehealth.com/pages/hume-bod… Latte: https://longevitylatte.shop TIMESTAMPS 0:00 — Cold Open 1:41 — Sponsor: Hume Body Pod 2:53 — Intro: The Cholesterol Paradox 4:48 — How Statins Work 5:36 — PCSK9 Targeting 7:38 — How VERVE-102 Works 9:16 — The Results 12:16 — Other Drugs and Pricing 15:12 — Natural Alternatives 17:42 — Other One and Done Drugs 18:40 — Longevity Latte SOURCES & LINKS NEJM paper (Vafai, Täubel, Patel, Kathiresan et al.): https://www.nejm.org/doi/full/10.1056… Eli Lilly press release on Phase 1 Heart-2 data: https://www.prnewswire.com/news-relea… ClinicalTrials.gov Heart-2 trial entry (NCT06164730): https://clinicaltrials.gov/study/NCT0… Verve Therapeutics FDA Fast Track designation announcement: https://vervetx.gcs-web.com/news-rele… Cohen and Hobbs 2006 NEJM paper (the foundational PCSK9 loss-of-function discovery): https://www.nejm.org/doi/abs/10.1056/.… Dave Ricks (Eli Lilly CEO) on Cheeky Pint with Patrick and John Collison: • Dave Ricks, CEO of Eli Lilly, on GLP-1s an… FOURIER trial (evolocumab cardiovascular outcomes): https://www.nejm.org/doi/full/10.1056… ODYSSEY OUTCOMES trial (alirocumab cardiovascular outcomes): https://www.nejm.org/doi/full/10.1056… PCSK9 LoF and diabetes (Mendelian randomization, Lancet Diabetes & Endocrinology): https://www.thelancet.com/journals/la… PCSK9 inhibition and diabetes risk review: https://pmc.ncbi.nlm.nih.gov/articles… StatPearls overview of PCSK9 inhibitors: https://www.ncbi.nlm.nih.gov/books/NB… Sardinia cholesterol paradox study: https://pmc.ncbi.nlm.nih.gov/articles… Statin pleiotropic effects review (mevalonate pathway): https://www.ncbi.nlm.nih.gov/pmc/arti… Berberine as a nature-made PCSK9 inhibitor review: https://www.ncbi.nlm.nih.gov/pmc/arti… Berberine for dyslipidaemias meta-analysis: https://pubmed.ncbi.nlm.nih.gov/30466… Pomegranate juice, carotid IMT, and LDL oxidation (Aviram 3-year study): https://www.clinicalnutritionjournal… LATEST EXCLUSIVE INTERVIEW: [Ariel Garten / Muse headset interview YouTube URL] ABOUT LONGEVITY SCIENCE NEWS Longevity Science News covers the latest breakthroughs in anti-aging research, regenerative medicine, longevity biotech, and the science of extending human healthspan and lifespan. Hosted by Emmett Short. Disclaimer: This video is for educational and informational purposes only and does not constitute medical advice. Consult a qualified clinician before making health or treatment decisions. EXCLUSIVE INTERVIEWS & BONUS CONTENT Patreon: https://patreon.com/u29506604?utm_med… YT Membership:
/ @longevitysciencenews PRODUCTION CREDITS Executive Producer – Keith Comito Host, Producer, Writer – Emmett Short.
Longevity Latte:
https://longevitylatte.shop.
TIMESTAMPS
0:00 — Cold Open.
1:41 — Sponsor: Hume Body Pod.
2:53 — Intro: The Cholesterol Paradox.
4:48 — How Statins Work.
5:36 — PCSK9 Targeting.
7:38 — How VERVE-102 Works.
9:16 — The Results.
12:16 — Other Drugs and Pricing.
15:12 — Natural Alternatives.
17:42 — Other One and Done Drugs.
18:40 — Longevity Latte.
SOURCES \& LINKS
NEJM paper (Vafai, Täubel, Patel, Kathiresan et al.): https://www.nejm.org/doi/full/10.1056…
Eli Lilly press release on Phase 1 Heart-2 data: https://www.prnewswire.com/news-relea…
ClinicalTrials.gov Heart-2 trial entry (NCT06164730): https://clinicaltrials.gov/study/NCT0…
Verve Therapeutics FDA Fast Track designation announcement: https://vervetx.gcs-web.com/news-rele…
Cohen and Hobbs 2006 NEJM paper (the foundational PCSK9 loss-of-function discovery): https://www.nejm.org/doi/abs/10.1056/.…
Dave Ricks (Eli Lilly CEO) on Cheeky Pint with Patrick and John Collison: • Dave Ricks, CEO of Eli Lilly, on GLP-1s an…
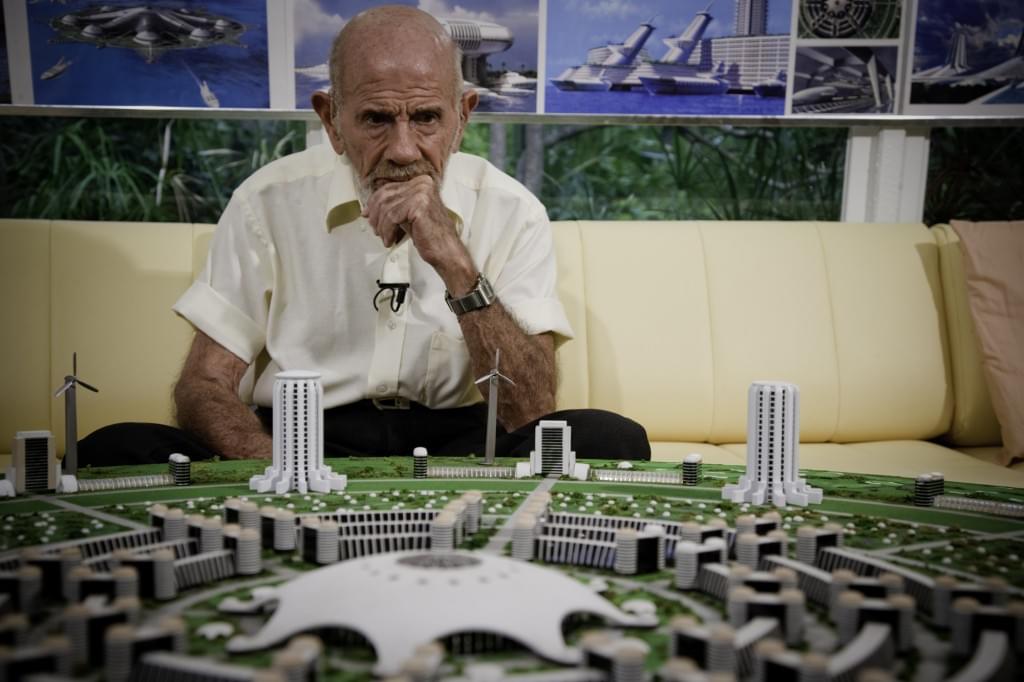

I have to confess something about this interview.
I really liked Jacque Fresco. Not as a thinker I was supposed to admire, but as a person: the humor, the humility, the scientific curiosity still burning at 97.
That made the disagreements harder, not easier.
Fresco spent almost a century arguing one idea. We apply the methods of #science to engineering, to medicine, to flight. Then we run our economies and our politics on opinion, tradition, and the preferences of the financial elite.
He thought we had it exactly inverted. Rigor for the machines, guesswork for the humans.
“Technology was never the hard part. The harder question is what kind of society we want it to serve.”
When doctors and scientists want to see inside a body, magnetic resonance imaging (MRI) is a powerful tool. MRI can noninvasively capture detailed images of the body’s muscles, organs, and bones. It can monitor blood flow to generate a map of brain activity. And with new sensors developed by bioengineers at MIT, MRI can track the kinds of molecules that make our brains and bodies work.
In the May 13 issue of the journal Nature Biomedical Engineering, a team led by Alan Jasanoff, the Eugene McDermott Professor in the Brain Sciences and Human Behavior at MIT, reports on their new sensors, which can brighten or dim MRI signals in response to specific molecular targets. The probes are designed to amplify the effect that each target molecule has on MRI signal, dramatically improving sensitivity over previous small-molecule sensors.
Jasanoff, who is also an associate investigator at the McGovern Institute for Brain Research, says the approach his team used should enable the development of MRI sensors that detect neurotransmitters and other important molecules in the brain.

Nine years after wowing the audience at the American Society of Clinical Oncology annual meeting with a CAR-T candidate that would become Carvykti—now the world’s most successful cell therapy—Legend Biotech’s scientific founder, Frank Fan, M.D., Ph.D., is returning to the spotlight with an entirely new playbook.
This time, Fan isn’t showcasing an autologous product engineered with each individual patient’s cells. Instead, with his new venture, Wondercel Therapeutics, Fan hopes an off-the-shelf universal CAR-T platform can tackle two bottlenecks of the cell therapy industry: massive production scalability and the pitfalls associated with gene editing.
“If this approach proves successful, the critical thing is that we can achieve linear scalability in CAR-T production capacity that can match traditional biologics,” Fan said in an interview with Fierce.