Thirdly, more recent approaches have begun to leverage deep learning (DL) methods. DL models such as U-Net12 have provided solutions for many image analysis challenges. However, they require ground truth to be generated for training. DL-based methods for SST cell segmentation include GeneSegNet13 and SCS14, though supervision is still required in the form of initial cell labels or based on hard-coded rules. Further limitations of existing methods encountered during our benchmarking, such as lengthy code runtimes, are included in Supplementary Table 1. The self-supervised learning (SSL) paradigm can provide a solution to overcome the requirement of annotations. While SSL-based methods have shown promise for other imaging modalities15,16, direct application to SST images remains challenging. SST data are considerably different from other cellular imaging modalities and natural images (e.g., regular RGB images), as they typically contain hundreds of channels, and there is a lack of clear visual cues that indicate cell boundaries. This creates new challenges such as (i) accurately delineating cohesive masks for cells in densely-packed regions, (ii) handling high sparsity within gene channels, and (iii) addressing the lack of contrast for cell instances.
While these morphological and DL-based approaches have shown promise, they have not fully exploited the high-dimensional expression information contained within SST data. It has become increasingly clear that relying solely on imaging information may not be sufficient to accurately segment cells. There is growing interest in leveraging large, well-annotated scRNA-seq datasets17, as exemplified by JSTA18, which proposed a joint cell segmentation and cell type annotation strategy. While much of the literature has emphasised the importance of accounting for biological information such as transcriptional composition, cell type, and cell morphology, the impact of incorporating such information into segmentation approaches remains to be fully understood.
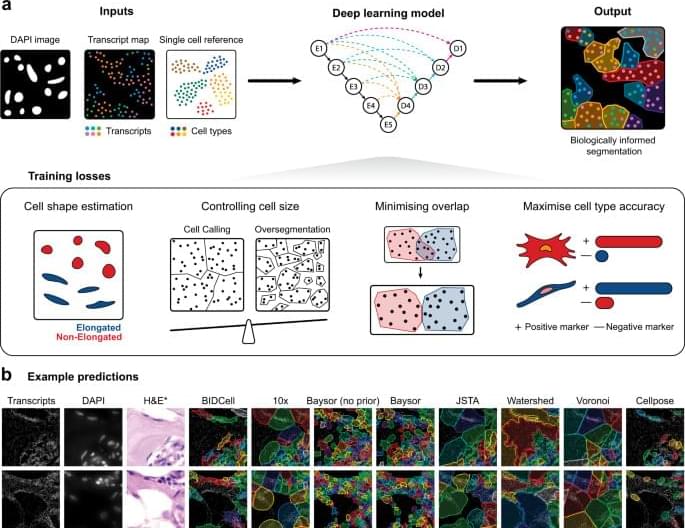
Here, we present a biologically-informed deep learning-based cell segmentation (BIDCell) framework (Fig. 1 a), that addresses the challenges of cell body segmentation in SST images through key innovations in the framework and learning strategies. We introduce (a) biologically-informed loss functions with multiple synergistic components; and (b) explicitly incorporate prior knowledge from single-cell sequencing data to enable the estimation of different cell shapes. The combination of our losses and use of existing scRNA-seq data in supplement to subcellular imaging data improves performance, and BIDCell is generalisable across different SST platforms. Along with the development of our segmentation method, we created a comprehensive evaluation framework for cell segmentation, CellSPA, that assesses five complementary categories of criteria for identifying the optimal segmentation strategies. This framework aims to promote the adoption of new segmentation methods for novel biotechnological data.