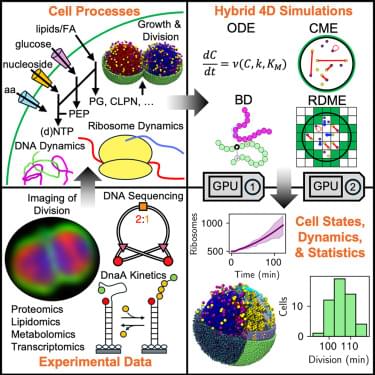
David S. Yu & team now show SIRT2 deacetylates MRE11 facilitating DNA binding to promote DNA end resection and ATM-dependent DNA damage signaling:
The figure shows MRE11 K393 deacetylation by SIRT2 promotes DNA end resection after ionizing radiation exposure, in the osteosarcoma cell line U20S.
1Department of Radiation Oncology and Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
2Department of Biology, Clark Atlanta University, Atlanta, Georgia, USA.
3Department of Biochemistry, Emory University School of Medicine, Atlanta, Georgia, USA.